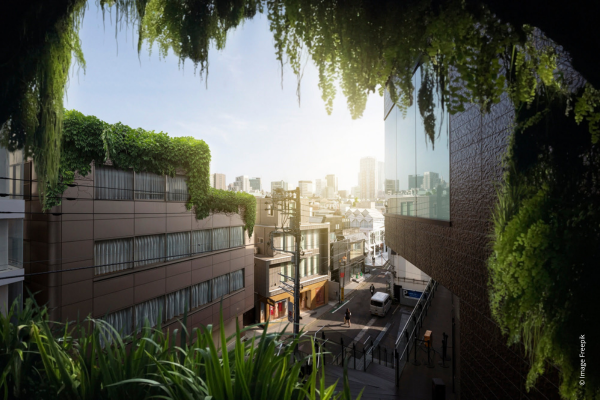
- Lire la suiteVisite PEPR : explorer des solutions durables face aux changements globauxEvènement / Annonce
Le 5 avril, le PEPR SOLU-BIOD porté par Xavier le Roux a été inauguré à Lyon en présence de personnalités scientifiques et politiques. Antoine Petit, PDG du CNRS, qui était présent à cette occasion, a visité en marge l'événement le laboratoire LEHNA, impliqué dans plusieurs PEPR visant à proposer des solutions durables sur les territoires, en réponse aux changements globaux.
- Lire la suiteCordées de la Réussite : des collégiens et lycéens accueillis tout au long de l'année à l'UCBLEvènement / Annonce
Véritable levier pour permettre aux collégiens et lycéens d'être accompagnés dans leur orientation, l'Université Claude Bernard Lyon 1 s'est totalement investie dans les Cordées de la Réussite.
- Lire la suiteConnaissez-vous la Cellule congrès de l’UCBL ?Colloque / Séminaire
Dans le cadre des missions de diffusion et de partage des savoirs, la Cellule Congrès de l’Université Claude Bernard Lyon 1 propose ses services et un accompagnement de proximité pour l’organisation d’événements scientifiques nationaux ou internationaux depuis 20 ans.
- Lire la suiteLa mission égalité-diversité organise son Festival hors de genre 2024Evènement / Annoncedu 7 mai 2024 au 24 mai 2024
Comme tous les ans à l'occasion de la journée internationale de lutte contre les haines anti-LGBTQI+, la mission égalité-diversité organise le festival Hors de genre qui favorise les représentations et interroge les normes de genre aux travers d'événements culturels, de formations et de rencontres.
AGENDA
- 29avr. 31maiSpectacle, Exposition Festival Les Arthémiades
- 07maiSpectacle Spectacle Drag & Talk (dans le cadre des Arthémiades)
- 16maiAction, Formation pro Formation webinaire « Accompagnement des élèves LGBTQI » avec Morgan N. Lucas
- 16maiAtelier / rencontre Carte blanche à Valérie Martinez
Hall de la BU
L'UNIVERSITÉ EN CHIFFRES
46 000 étudiantes et étudiants
450 diplômes & 10 000 diplômés par an
7 523 publications en 2022
13,5 M€ dédiés au soutien direct à la recherche en 2023 (fonctionnement, appels à projets)
1ʳᵉ université française en nombre de brevets déposés (INPI 2022)
LES GRANDS PROJETS
Les grands projets de l’Université s’inscrivent dans sa stratégie académique et ses valeurs de diversité, d’agilité, de solidarité et de responsabilité. Ces projets structurants, alignés sur les priorités de l’État, renforcent ses missions de formation, de recherche et d’innovation de rang mondial et permettent le développement d’infrastructures et de services conformes aux standards internationaux.
Ils façonnent ainsi une université toujours plus ancrée sur son territoire et résolument tournée vers l’avenir. Par ces projets variés, l’Université Claude Bernard Lyon 1, avec ses nombreux partenaires, abordent, via une approche pluridisciplinaire, les défis complexes de notre société : transformation numérique, inclusion, environnement, intelligence artificielle, santé…
Les projets scientifiques confirment le rôle moteur de l’Université pour le développement d’une recherche d’excellence et de l’innovation scientifique et affirment sa place d’acteur engagé pour un monde plus soutenable et durable.
INCLUSION & NUMÉRIQUE - Démonstrateur INCLUDE
RECHERCHE & FORMATION EN SANTÉ - SHAPE-Med@LYON
FORMATION PAR LA RECHERCHE - Projet Graduate +
NUMÉRIQUE - Centre de Calculs et de Données LyonTech - la Doua
SCIENCES POUR TOUS
SCIENCES EN RÉCITS
 Découvertes, prix, innovations : à l’Université Lyon 1, des parcours et des trajectoires extraordinaires se dessinent chaque jour. Comment ces aventures sont-elles vécues par leurs protagonistes ? Ils sont étudiants, étudiantes, enseignants, enseignantes, scientifiques, et témoignent en quelques minutes des temps forts de leurs histoires hors du commun.
Découvertes, prix, innovations : à l’Université Lyon 1, des parcours et des trajectoires extraordinaires se dessinent chaque jour. Comment ces aventures sont-elles vécues par leurs protagonistes ? Ils sont étudiants, étudiantes, enseignants, enseignantes, scientifiques, et témoignent en quelques minutes des temps forts de leurs histoires hors du commun.Entrez dans les coulisses d’une Université de sciences, technologies, santé et sport, et écoutez les récits de vie de celles et ceux qui l’animent.
UNIVERSITÉ OUVERTE
 Découvrir la biologie ou la physique, s’initier à l’histoire de l’art et des sciences, s’émerveiller de la beauté de notre monde et s’interroger sur celui que l’on bâtit pour demain… C’est la mission de L'Université Ouverte de partager avec le plus grand nombre les progrès scientifiques et les questions qu’ils posent.
Découvrir la biologie ou la physique, s’initier à l’histoire de l’art et des sciences, s’émerveiller de la beauté de notre monde et s’interroger sur celui que l’on bâtit pour demain… C’est la mission de L'Université Ouverte de partager avec le plus grand nombre les progrès scientifiques et les questions qu’ils posent.Près de 190 conférences et ateliers pour apprendre, approfondir vos connaissances mais aussi forger votre propre opinion sur des sujets à fort enjeu sociétal ou à controverse.



 fr
fr en
en
 es
es